While Sanger sequencing is limited to a single DNA fragment, high throughput sequencing allows the simultaneous acquisition of data related to millions of DNA fragments in a so-called library.
As a highly scalable technology, Next generation sequencing can be used to analyse the genetic material of any organism:
- Sequencing of genomes still not referenced (De novo Sequencing)
- Re-sequencing of referenced genomes in order to find new genetic variants
- Analysis of transcripts: RNA-Seq
- Identification of point mutations (SNPs), insertions or deletions within DNA fragments
- Detection of different organisms (biodiversity) present in a given sample (analysis of the intestinal microbiome for example) by the analysis of target sequences
- Determination of DNA methylation sites
- Study of DNA/RNA - protein interactions...
There are several NGS sequencing technologies available depending on the manufacturer:
- 454 technology or pyrosequencing by Roche
- Ligation sequencing by Applied Biosystems
- Sequencing by synthesis by Illumina

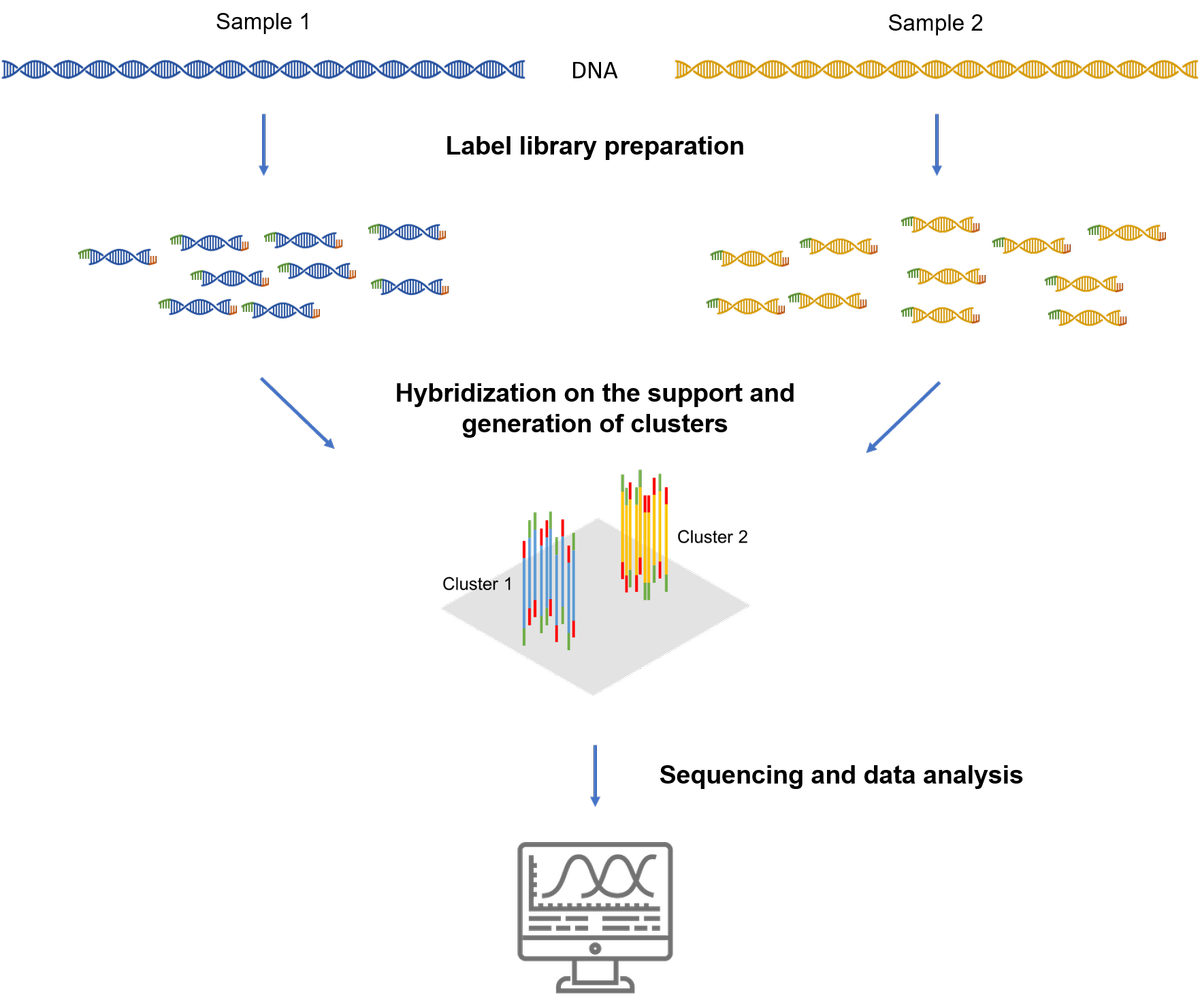
Illumina technology is based on the sequencing, base after base of clusters. Clusters are groups of identical DNA fragments positioned close to each other on a solid support.
Sequencing on Illumina equipment is carried out/performed as follows:
- Preparation of library(ies): it consists, either, in a chemical, enzymatic or mechanical lysis of the DNA (cDNA or gDNA) into fragments of reduced and homogeneous size at the ends of which small specific sequences called labels (tags) will be added, or in the targeted amplification of fragments using complementary primers comprising the labels. More informations on the preparation of librairies.
- Amplification by PCR of each of the components of the library from the the sequences complementary to sequences of the labels fixed on the solid support to generate the clusters. Each cluster contains between 1,000 and 5,000 identical molecules. According to the support (flow cell) used, several tens of millions of clusters can form on the support.
- Determination of the sequence by incorporation of fluorescent dNTPs. Reading may be done base by base from a label. It is also possible to read the fragments in "Paired-end read", that means in both directions from both lables.
NGS sequencing results are characterized by two important notions:
- Coverage: percentage of the library that is effectively sequenced
- Depth: (average) number of times each base is read
These quantities concepts are generally defined upstream of the experiment and are have to be modulated according to the project. For example, for De novo sequencing, high coverage and depth will be recommended, while for amplicon sequencing, a high depth will be preferred.

BIOMNIGENE assists you in your high throughput sequencing projects.
Trained by Illumina, the world leader in sequencing, our engineers will offer you the experimental approach that fits your objectives. Our laboratory is equipped with two Illumina second-generation devices: the MiSeq and the iSeq100 which allow microorganism genome and transcriptome, microbiome study, animal genotyping, and other applications ...
Collected raw data as well as a report of the experiment results will be returned to you in FastQ format on a USB key.
(Some) bioinformatics analyses can be performed by BIOMNIGENE, such as the determination of variants or the characterization of a microbiome. For further analysis, we will redirect you to one towards our partners.



