You design innovative immunotherapies to fight against certain diseases.
Your years of research and development have allowed you to develop a cell line that produces an antibody specifically efficient to fight against a given disease.
We are able to determine the nucleotide sequences encoding the different chains of this antibody, allowing you to conserve them and reproduce the molecule of interest, even if your cell line was to disappear.


The in-house development of two methodologies allows us to respond to all requests regarding antibody sequencing such as :
- Sequencing of the variable regions of light and heavy chains
- Determination of the sequence of your antibody "leader” region
- Complete sequencing of the constant regions of the heavy and / or light chains
- Analysis of cells producing polyclonal antibodies ...
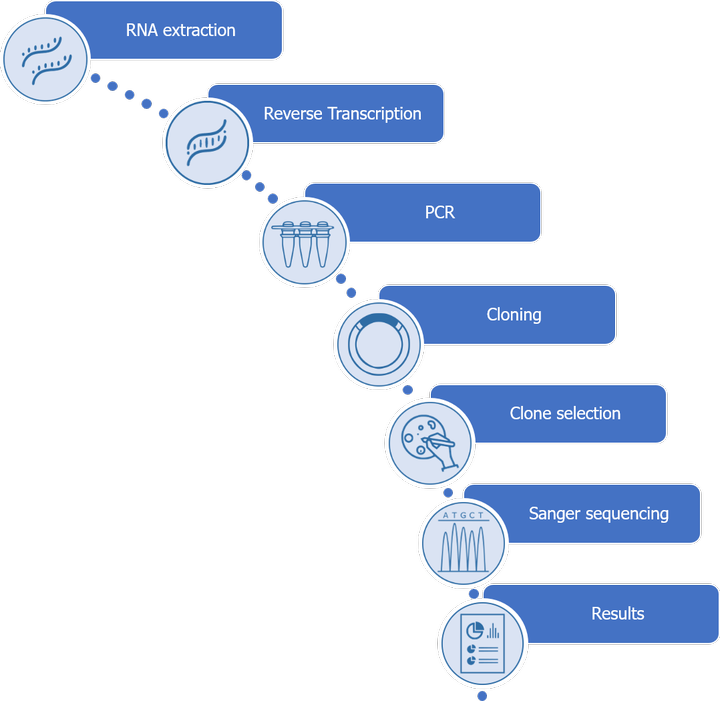
From frozen cells, we first extract total RNAs on which an RT-PCR is performed. This reaction allows us to obtain cDNA fragments that will be amplified and then cloned after insertion in a cloning vector.
Bacteria transformed with the recombinant plasmids generated will be screened so as to sequence the inserts by the Sanger method.
Sequences must be checked on several clones in order to be validated. The number of validations depends on the number of clones obtained.
A report containing the different sequences found in the cultures is sent to you.
The report provided contains according to your requests:
- The annotated nucleotide sequences of the heavy and light chain domains depending on your need
- The corresponding protein sequences
- A theoretical confirmation of the productivity of the heavy or light chain by bioinformatics
- If applicable, other productive nucleotide and protein sequences determined by sequencing the bacterial clones
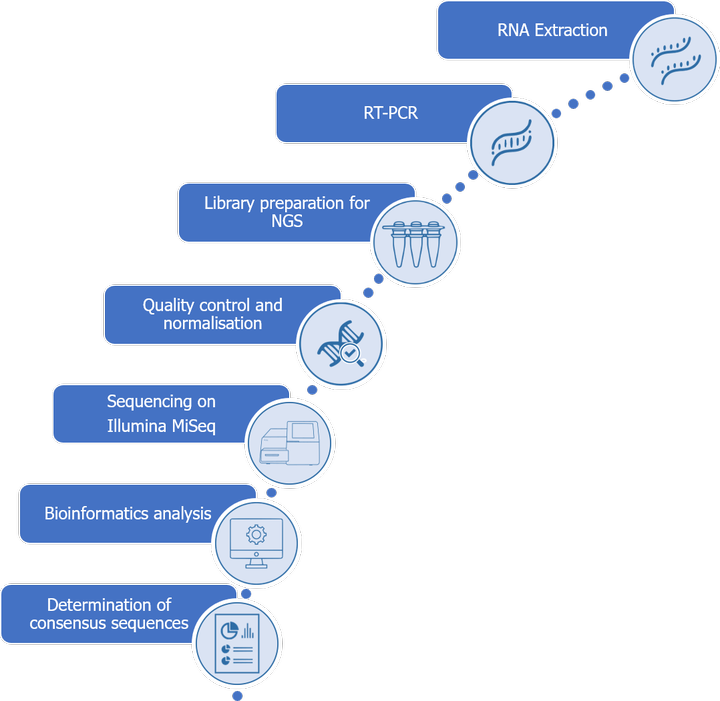
Total RNA is first extracted from frozen cells and an RT-PCR is performed.
The obtained cDNAs are then processed to obtain an NGS library, quality controls are performed in order to check the quantity of material and the length of the fragments composing the library.
A normalization of the libraries is carried out in order to standardize the quantities deposited on the flow-cell.
After sequencing on an Illumina MiSeq device, the data is bioinformatically processed to determine the different sequences expected.
The report provided contains according to your requests:
- The annotated nucleotide sequences of/from the heavy and light chain domains, according to your need
- The read depth reflecting the degree of reliability of each base
- The corresponding protein sequences
- A confirmation of the productivity of the heavy or light chain by bioinformatics.
- If applicable, other productive nucleotide and protein sequences
Our methodologies allow us to meet all your antibody sequencing needs, from single hybridoma sequencing to large series. We can also analyze polyclonal antibody cultures.
Sequencing by cloning
Our standard services allow you to obtain the nucleotide sequences of your variable, heavy and light domains.
We offer, on request, other options such as:
- The Leader Sequence
- The constant domains of the heavy chain (CH1, CH2, CH3)
- The constant domain of the light chain (CL)
NGS sequencing
Our high-throughput approach to antibody sequencing allows you to sequence from 24 to 384 hybridomas in a single assay.
For more information and personalised pricing, please contact us by e-mail or by telephone using the details on our contact page.



